560
Member
Elucidation of biological functions using multilevel techniques to evaluate cardiovascular functions and its clinical application
Our cardiocirculatory function is mainly controlled by muscular organs composed of striated muscles (heart and skeletal muscles) and smooth muscle (blood vessels). Our group aims to elucidate the molecular mechanisms underlying transition of the muscles from adaptation to maladaptation against various stress (hemodynamic load and environmental stress) using multi-level techniques to evaluate cardiovascular functions (in vivo and in vitro), and work toward practical application (e.g., drug discovery and fostering). In particular, we are focusing on mitochondria, energy-producing organs, and investigating the mechanism of muscle repair and regeneration from the viewpoint of mitochondrial quality control. We aim to develop a novel therapeutic strategy for refractory diseases.
Disruption of redox (reduction/oxidation) dynamics is closely related to the onset of various diseases including cardiocirculatory diseases. We are focusing on highly reactive sulfur metabolites (supersulfides) and conducting sulfur redox biology for cardiovascular homeostasis and diseases. In addition, we address the inclusive research to elucidate the mechanism underlying maintenance and transfiguration of cardiocirculatory homeostasis via multi-organ interactions by combining non-invasive measuring methodologies of motor functions and those cardiovascular functions. Our laboratory has various techniques and equipment to drive the above researches.
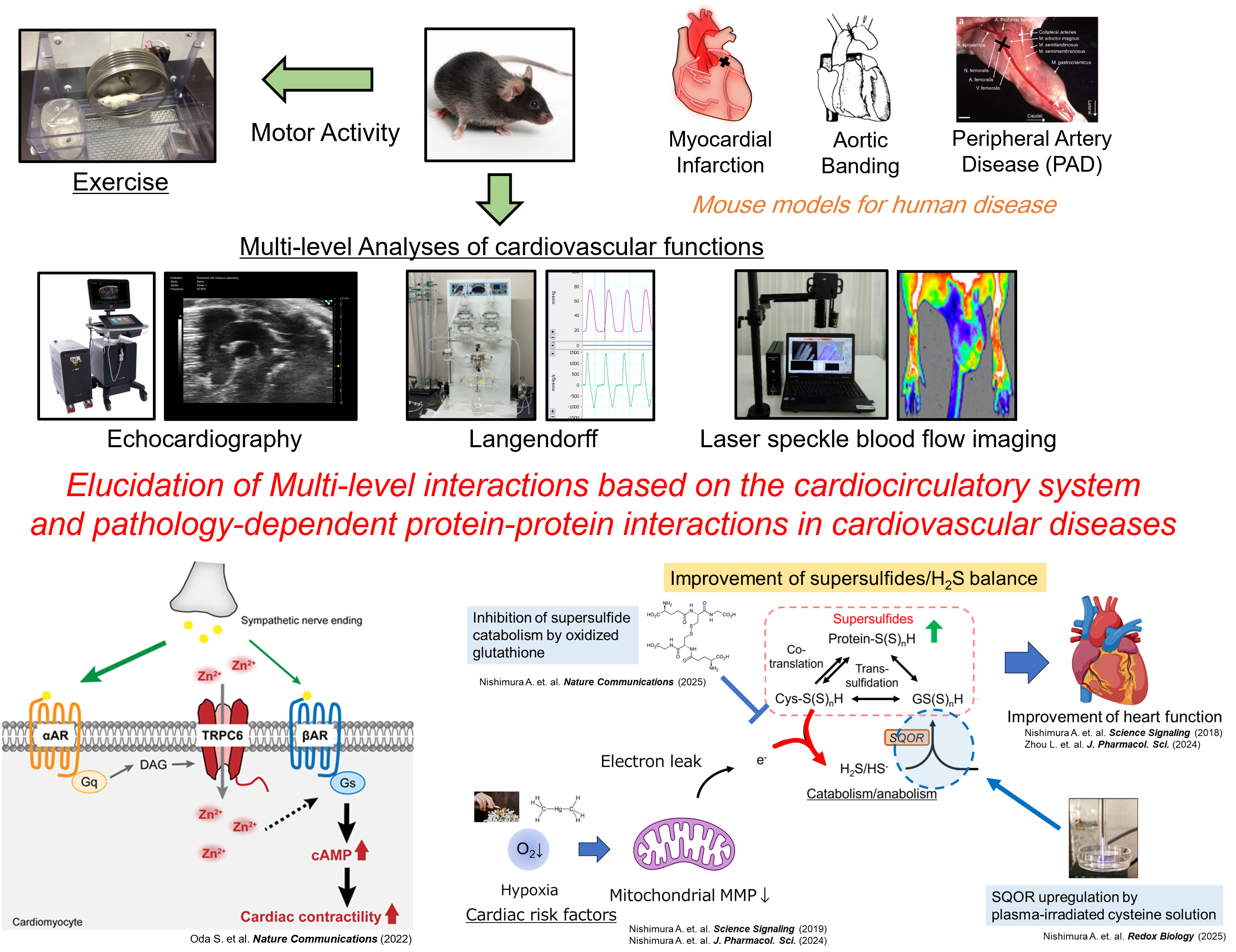
Figure. Measuring systems for cardiovascular functions and summary of our research using these systems
Selected publications
*A. Nishimura et al. Nat. Commun. 16. 276 (2025)
*A. Nishimura et al. Redox Biol. 79, 103445 (2025)
*L. Zhou et al. J. Pharmacol. Sci. 155. 121-130 (2024)
*A. Nishimura et al. J. Pharmacol. Sci. 154. 127-135 (2024)
*X. Tang et al. Mar. Drugs. 21. 52 (2023)
*S. Oda et al. Nat. Commun. 13. 6374 (2022)
