Giant viruses are group of extremely large viruses, some of which are larger than typical bacteria. They have extremely large genomes and contain many unique genes not found in other living organisms. Here, a research team consisting of Kyoto University, Tokyo University of Science, National Institute for Physiological Sciences, and Tokyo Institute of Technology, has discovered a new giant virus infecting Amoeba from a hot spring water in Japan. It’s genomic, structural, and cytological features were studied in detail.
Based on the encystment affect the virus provides to the host amoeba, the new giant virus was named "Medusavirus". Genomic and structural features indicate that Medusavirus is distantly related with other giant viruses, and we have concluded that it belongs to a new taxonomical "family" of giant viruses, “Medusaviridae”. One major genomic feature of Medusavirus is that it had the full set of histone proteins. Histones are eukaryotic proteins involved in the genomic DNA folding within the cell nucleus, and is composed of five essential proteins. Some viruses were already known to have part of them, but up until now, never the full five histone proteins. Some Eukaryotic DNA-related genes had been proposed to have derived from viruses. Our new results may also support this evolutionary hypothesis.
In order to unravel the long term coevolution history of eukaryotic cells and viruses in molecular level, we intend to study the infection process of Medusavirus in more detail, including the role of the viral histones. The current results will be published online in Journal of Virology on February 6th, 2019.
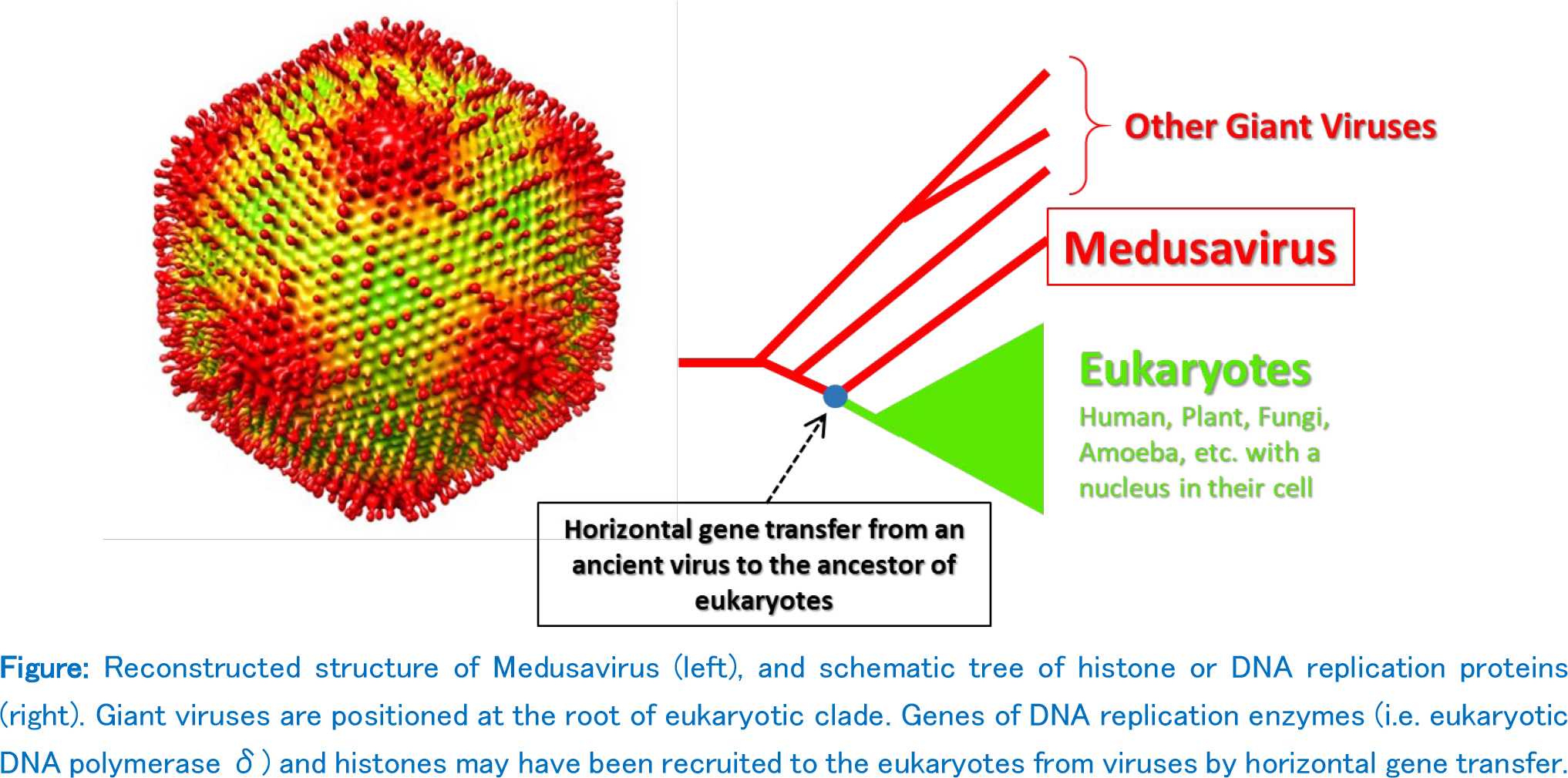
The research team isolated a new giant virus named "Medusavirus" from a hot spring water in Japan. Phylogenetic analysis of several key genes including DNA polymerase placed the genes from this new giant virus at the root of eukaryotic clades. And quite surprisingly, Medusavirus encoded all five histone genes, which was never found in a full set in any other viruses. In addition, many Medusavirus genes were also found from its host amoeba genome, including the major capsid protein. This indicate that indeed amoeba has been the genuine natural host of the Medusavirus, and lateral gene transfers repeatedly took place between the two from the ancient times. Altogether, our results suggest that Medusavirus is a unique giant virus that still preserves the ancient footprints of the virus-host evolutionary interactions, and thus provide clues to elucidate the long lasting coevolution between giant viruses and eukaryotes.
In the early 2000s, a novel huge virus was found, which completely flipped out understanding of viruses. The new virus infecting amoeba, a unicellular eukaryote (protist), was named “Mimivirus”. Surprisingly, the virion particle size and genome length of Mimivirus was larger than some single-celled organisms, and the general concept in biology as viruses being small and simple, was completely overturned. After the discovery of Mimivirus, researchers from all over the world began hunting for even larger giant viruses, and Pandoravirus, Pithovirus, Marseillevirus and so on was discovered. Recently in Japan, group of Prof. Masaharu Takemura of Tokyo University of Science has discovered Tokyovirus (a member of the Marseillevirus), Mimivirus shirakomae (a member of the Mimivirus), and others.
Eventually, the more you begin to know about these giant viruses, you will be awed by their surprisingly vivid and complex life-like systems. And at the end, we end up having fundamental questions such as "Are viruses alive?", and also a more challenging and provocative hypothesis such as "Did these viruses evolved from cells?", "Is DNA a viral invention?", and "Is the cell nucleus derived from viruses?" have come to be proposed.
Here, the collaborative research team has isolated a new giant virus from a sediment containing hot water sample from a hot spring region in Hokkaido, Japan. The virus was isolated using an amoeba co-culture system, and its infection process, particle structure and genome composition was studied in detail. The results indicate that this giant virus is different in various properties from the other known giant viruses.
 The host of the new giant virus is amoeba, and the viral infection results in some amoeba cells to form thick surface film and become dormant (encystment). This reminds the Greek mythical monster "Medusa", capable of transforming onlookers into a stone, and the research team named this new virus "Medusavirus".
The host of the new giant virus is amoeba, and the viral infection results in some amoeba cells to form thick surface film and become dormant (encystment). This reminds the Greek mythical monster "Medusa", capable of transforming onlookers into a stone, and the research team named this new virus "Medusavirus".
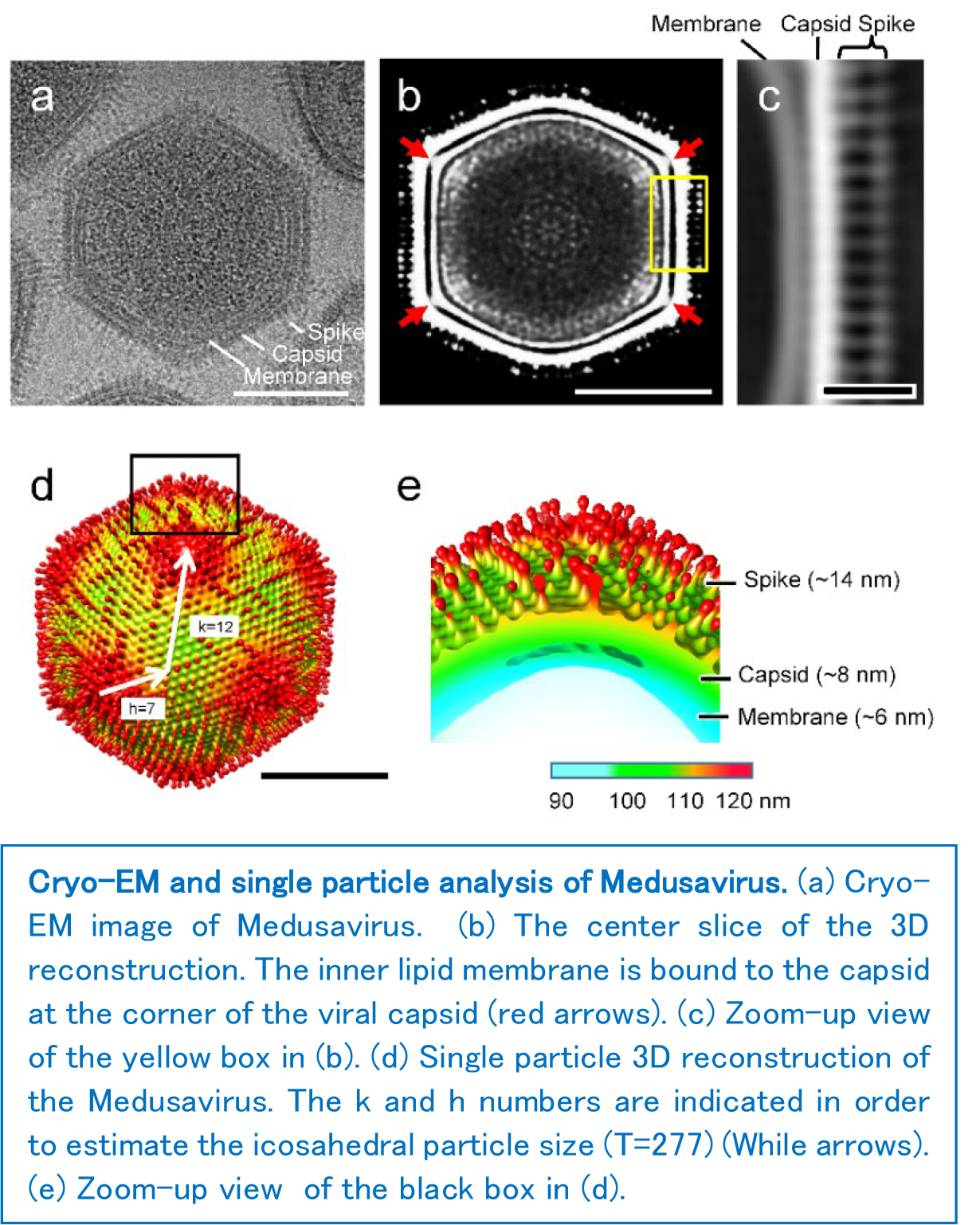
Among the other known giant viruses, Medusavirus is relatively small, with a particle size of 260 nm and a genome length of 380k base pairs. However, cryo-electron microscopy (cryo-EM) and single particle analysis revealed a unique particle morphology, where the viral capsid surface is covered with spikes with spherical heads. Its genomic content was also characterized, and out of 461 total encoding proteins, 61% (279) were novel genes which had no homologous sequences in the public database. Microscopic observation of the infection process suggested that the replication of the viral genome is completed within the amoeba cell nucleus, which is different from the other giant viruses. These results, together with the molecular phylogenetic analysis, the research team concluded that Medusavirus represents a new viral family, Medusaviridae. In viral taxonomy, "family" is one of the highest classification level.
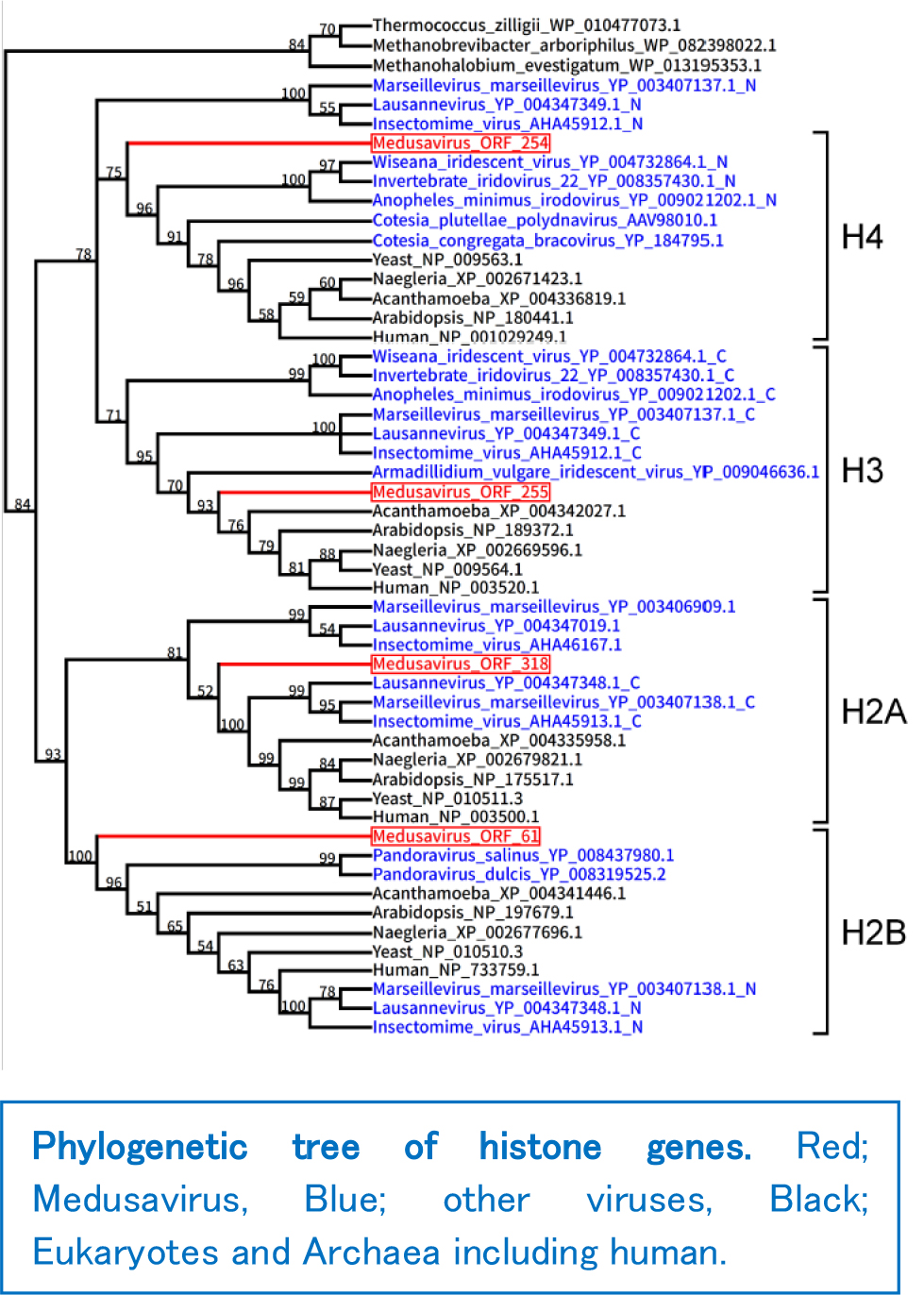 The most striking feature of the Medusavirus genome was that it encoded the whole set of five histone genes (H1, H2A, H2B, H3 and H4). Some other giant viruses such as Marseillevirus and Panduravirus were also known to encode part of histone genes, but Medusavirus is the first virus ever to be found with the complete set. The presence of viral encoded histone protein was also detected from the Medusavirus particle (virion). The result of molecular phylogenetic analysis was even more interesting. The phylogenic position of these histone genes branched out from the root of the modern eukaryotic clade. This suggests that their origin may be older than that of the eukaryotic common ancestors. At least, the results indicate that the viral histone genes did not derive from a particular extant eukaryotic lineage. One plausible scenario is that the ancestors of eukaryotes may have acquired histone genes from ancient viruses. Phylogenetic analysis of Medusavirus DNA replication enzyme gene also indicated similar evolutionary scenario.
The most striking feature of the Medusavirus genome was that it encoded the whole set of five histone genes (H1, H2A, H2B, H3 and H4). Some other giant viruses such as Marseillevirus and Panduravirus were also known to encode part of histone genes, but Medusavirus is the first virus ever to be found with the complete set. The presence of viral encoded histone protein was also detected from the Medusavirus particle (virion). The result of molecular phylogenetic analysis was even more interesting. The phylogenic position of these histone genes branched out from the root of the modern eukaryotic clade. This suggests that their origin may be older than that of the eukaryotic common ancestors. At least, the results indicate that the viral histone genes did not derive from a particular extant eukaryotic lineage. One plausible scenario is that the ancestors of eukaryotes may have acquired histone genes from ancient viruses. Phylogenetic analysis of Medusavirus DNA replication enzyme gene also indicated similar evolutionary scenario.
Furthermore, comparison of the amoeba and Medusavirus genome revealed that massive gene transfers (horizontal gene transfers) occurred between the two, in the course of evolution. The direction of gene transfers were in both directions, from ameba to virus, and from virus to ameba. Interestingly, one unusual gene that amoeba received from the virus was the major viral capsid protein gene, which makes the shell of the viral. Medusavirus may be exceptionally good at exchanging genes with its host.
In the future, the research team aims to elucidate the infection process of the Medusavirus in more detail, such as the role of viral histones, by using electron microscopy, transcriptome analysis, proteome analysis, biochemical analysis of viral proteins, etc. Through such studies, the research team wish to unravel how the billion years’ co-evolution occurred between giant viruses and eukaryotes.
Viruses are thought to have played an important role in the evolution and development of life. With the discovery of this Medusavirus, we were able to observe new traces of how the giant viruses got involved in the evolution of ancient eukaryotes. Of course it is indeed very interesting if viruses actually affected the birth of eukaryotes. The research team will continue further studies and deepen our understanding of the origins of viruses and life.
This study is the result of collaboration of experts in virus hunting, molecular biology, structural biology, bioinformatics. By collaborating with researchers from different fields, the research team was able to characterize the Medusavirus, which belonged to the new "Medusaviridae" family, fairly rapidly. The research team will continue this adventure exploring the attractive world of the virus.




Kyoto University
Tokyo University of Science
National Institute for Physiological Sciences(NIPS)
Tokyo Institute of Technology