Our state-of-the-art two-photon fluorescence lifetime imaging microscopes allow us to image protein activity and protein-protein interaction in living cells in deep tissue such as brain slice and brain of a living mouse. We accept the collaborative research using the fluorescence lifetime imaging microscope for imaging the activity and interaction of various signaling proteins. We also welcome students to pursue Ph.D. degrees, especially students who are interested in molecular imaging.
In addition to the cutting-edge microscope techniques, we try to develop novel fluorescent proteins and light-controllable signaling proteins. By far, we succeeded in visualizing the activities of signaling proteins in the dendritic spine of hippocampal neurons by using two-photon microscopy by combining the photo-activatable probes, new fluorescent proteins, electrophysiology. These techniques will enable us to reveal the system of neural networks and underlying molecular mechanisms in a living mouse neuron.
Our mission is to reveal “missing links” underlying molecular functions and physiological functions in a living body. We believe that the development & application of optical imaging methods will reveal the biological system at the cellular level.
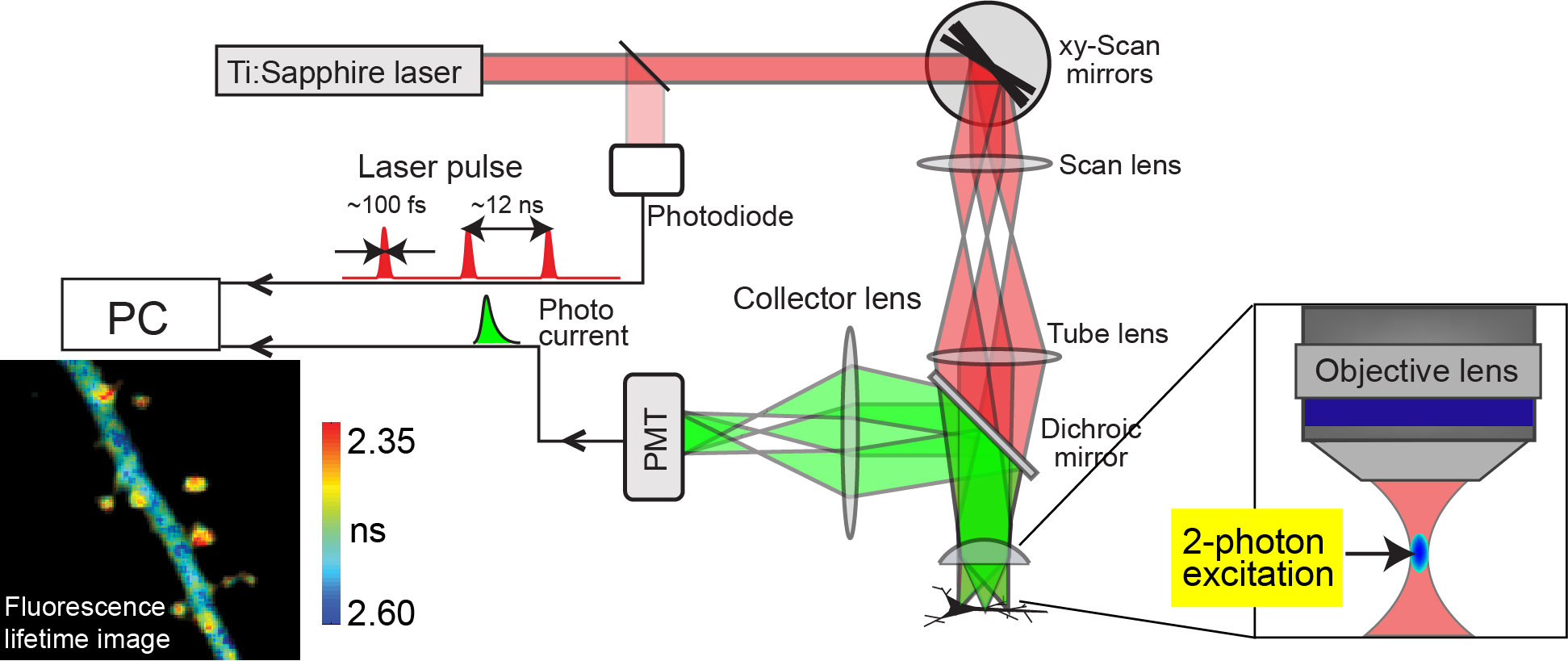
Figure 1.
Two-photon excitation is the phenomenon that two photons of half energy than needed for one photon excitation can excite a fluorescent molecule. The advantages of 2-photon excitation are 1) Because infrared light is used for excitation, it minimizes excitation-light scattering in the tissue 2) Because 2-photon excitation happens only at the focal point of an objective lens, the background signal is strongly suppressed. These effects enable us to image cells and subcellular structures in deep tissue with high spatial resolution. Recently, the combination of 2-photon excitation and fluorescence lifetime imaging method enabled us to image the protein-protein interaction or structural change of protein in deep tissue such as brain slice. The fluorescence lifetime is measured by counting the arrival time of signal photon at the detector upon a laser pulse. After making histogram of lifetimes at each pixel by repeating this measurement, the pixel-by-pixel lifetime image is constructed in a pseudocolor format.