|
A collaborative research team led by Kazuyoshi Murata determined the capsid structure of the giant Melbournevirus at 4.4 Å resolution using cryo-electron microscopy. By applying a block-based reconstruction method, they significantly improved 3D imaging resolution and revealed the detailed protein organization of the large capsid. These findings provide insights into how complex viral structures are built from limited protein components and are expected to advance understanding of viral evolution, infection mechanisms, and potential applications in drug delivery and material design. The study was published in Viruses on April 2, 2026. |
A research group led by Project Professor Kazuyoshi Murata at the Exploratory Research Center on Life and Living Systems (ExCELLS) / National Institute for Physiological Sciences (NIPS), in collaboration with Senior Researcher Kenta Okamoto at Uppsala University and Professor Chantal Abergel at Aix-Marseille University, has successfully determined, for the first time in the world, the capsid (outer shell) structure of Melbournevirus—a member of the giant virus family—at a resolution of 4.4 Å using cryo-electron microscopy (cryo-EM).
In this study, the researchers applied a “block-based reconstruction method” to the analysis of electron microscopy images, achieving a dramatic improvement in the resolution of the three-dimensional reconstruction (Figure 1). This enabled them to elucidate the detailed arrangement of proteins constituting the massive capsid (250 nm in diameter) (Figure 2).
These findings contribute to understanding the fundamental principles by which large, uniform, and robust structures are formed from a limited set of proteins. In addition, the results are expected to advance our knowledge of viral evolution and infection mechanisms, and may also be applied to the design of inclusion compounds and drug delivery carriers.
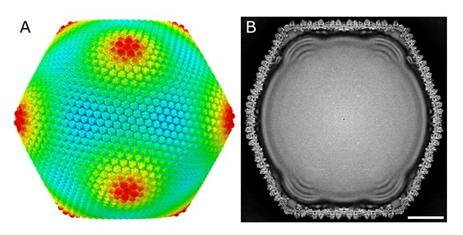
Figure 1. Overall structure of Melbournevirus at 4.4 Å resolution
Melbournevirus is an icosahedral giant virus with a diameter of approximately 250 nm. It is covered by capsid proteins composed of MCP and mCP, and it contains a folded DNA genome with characteristic protrusions inside. (A) External view. Colors indicate the distance from the particle center: blue, 1050 Å; green, 1150 Å; red, 1250 Å. (B) Cross-sectional view. The scale bar represents 500 nm.
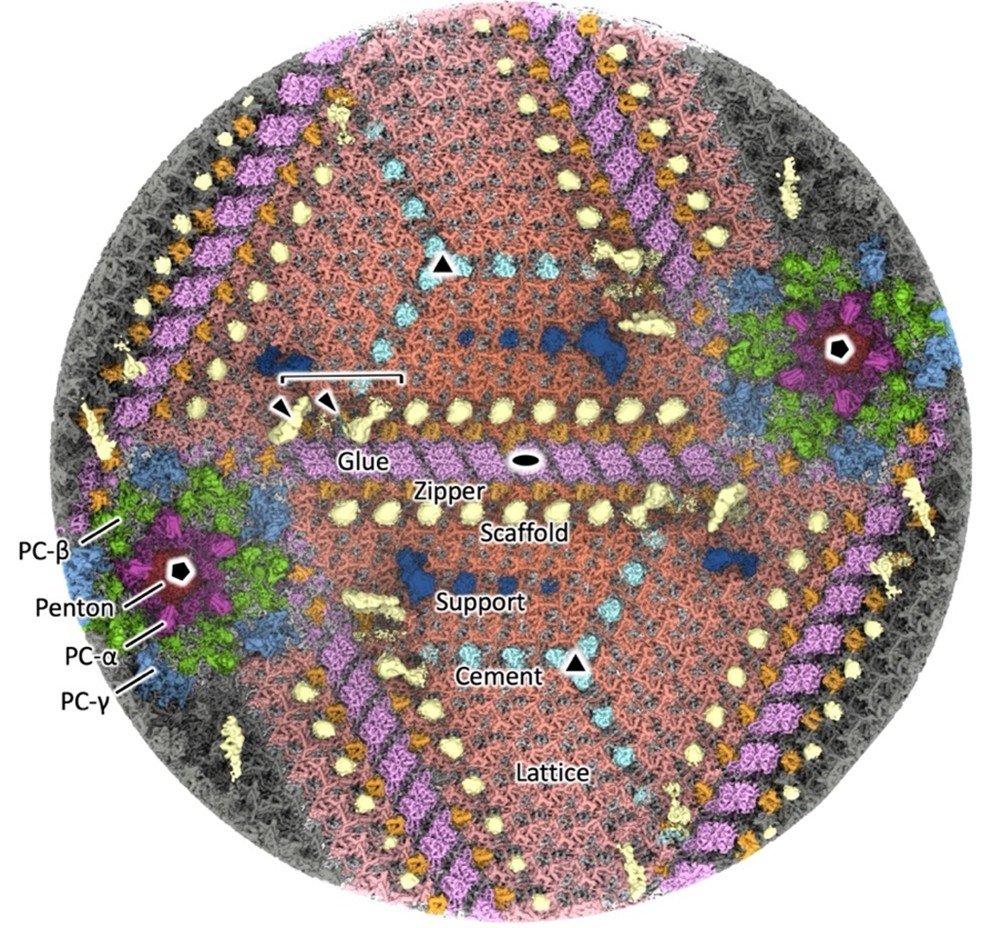
Figure 2. Interior view of the viral capsid
The 12 pentagonal vertices (two of which are shown in the diagram) are composed of protein components designated as Penton (black), PC-α (magenta), PC-β (green), and PC-γ (blue). The 20 triangular facets located between the vertices (two of which are shown in the diagram) consist of Cement (cyan), Lattice (vermillion), and Support (ultramarine) protein components, which are interconnected by Glue (purple) and Zipper (orange) protein components. On the inner side of the capsid, Scaffold (yellow) protein components are arranged along the junctions.
Funder
1. Ministry of Education, Culture, Sports, Science and Technology: JP19H04845
2. Japan Agency for Medical Research and Development: JP24ama121005, JP25ama121005
3. Swedish Research Council: 2018-03387
4. Swedish Foundation for International Cooperation in Research and Higher Education: JA2014-5721
5. Swedish Research Council for Environment, Agricultural Sciences, and Spatial Planning: 2018-00421
6. Royal Swedish Academy of Sciences: BS2018-0053
Journal
The 4.4 Å Capsid Structure of the Giant Melbournevirus Belonging to the Marseilleviridae Family
Raymond N. Burton-Smith, Chantal Abergel, Kenta Okamoto, Kazuyoshi Murata*(*Corresponding author)
Viruses(April 2, 2026.)
5116