Living organisms are built from a wide variety of biomolecules, including proteins, and their activities are sustained by the chemical reactions these molecules drive. In our research division, we use a technique called cryo-electron microscopy (cryo-EM) to investigate the structures of biomolecules and explore the fundamental question, “What is life?” at the molecular level.
Cryo-electron microscopy is a method in which biological samples are rapidly frozen and observed under an electron microscope while kept at very low temperatures. This approach allows us to analyze biomolecules at high resolution in a state that is close to their natural, living condition.
Our division is equipped with state-of-the-art instruments, including a 300 kV cryo-electron microscope (TITAN Krios G4) for high-resolution data collection, a 200 kV transmission cryo-electron microscope (JEM-2200FS) for sample screening, and a cryo-focused ion beam system (Aquilos 2) for preparing tomography samples. (Fig. 1, right).
The high-resolution images of proteins obtained from these instruments are processed using high-performance computers to reconstruct the three-dimensional structures of biomolecules.
Figure 2 shows the structure of amyloid fibrils determined by single-particle analysis using cryo-EM (Yagi-Utsumi et al., 2025). This result provides new insights into the process of amyloid fibril formation.
We welcome motivated young researchers and graduate students who are interested in structural biology to join our laboratory.
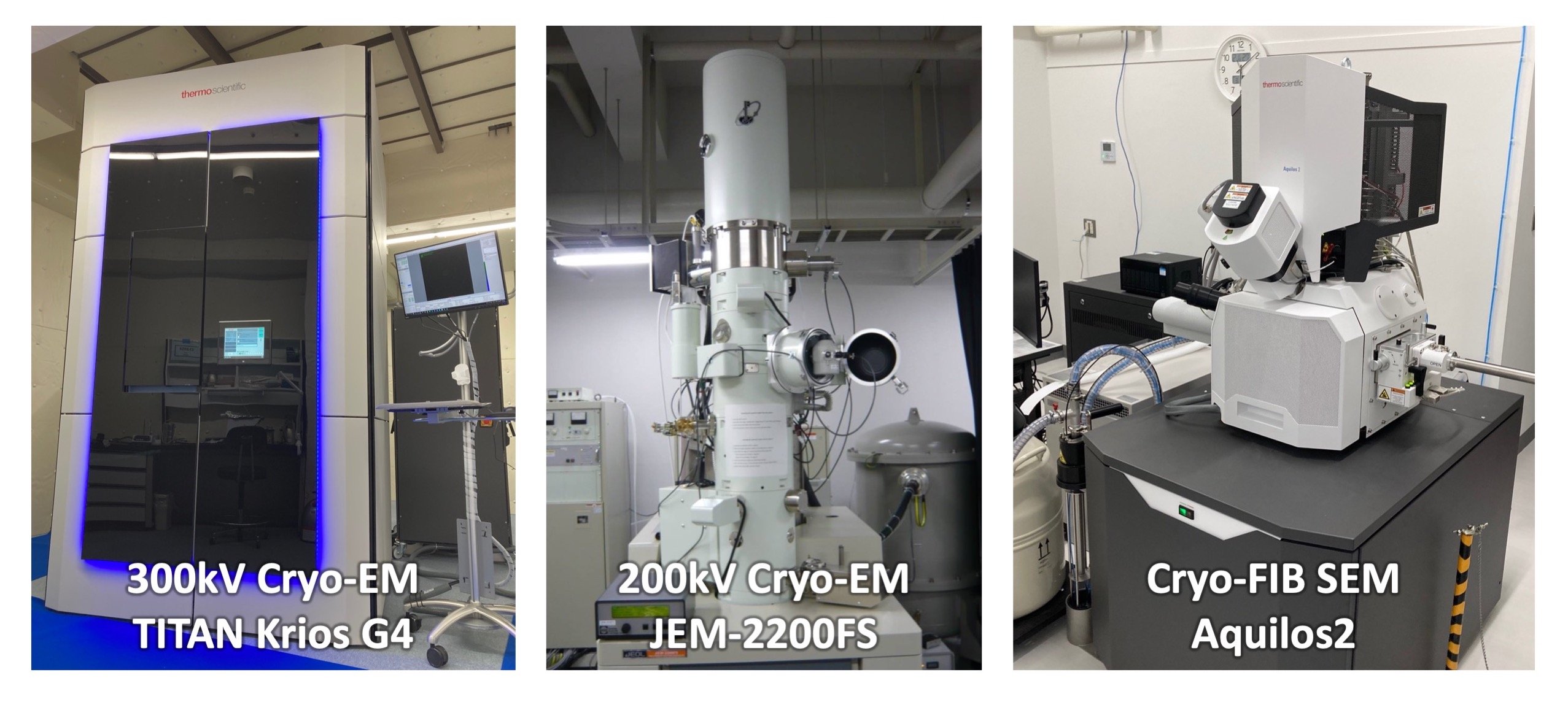

Fig. 1
300kV cryo-EM, TITAN Krios G4 (right), 200kV cryo-EM, JEM2200FS (middle), and cryo-FIB SEM (right).
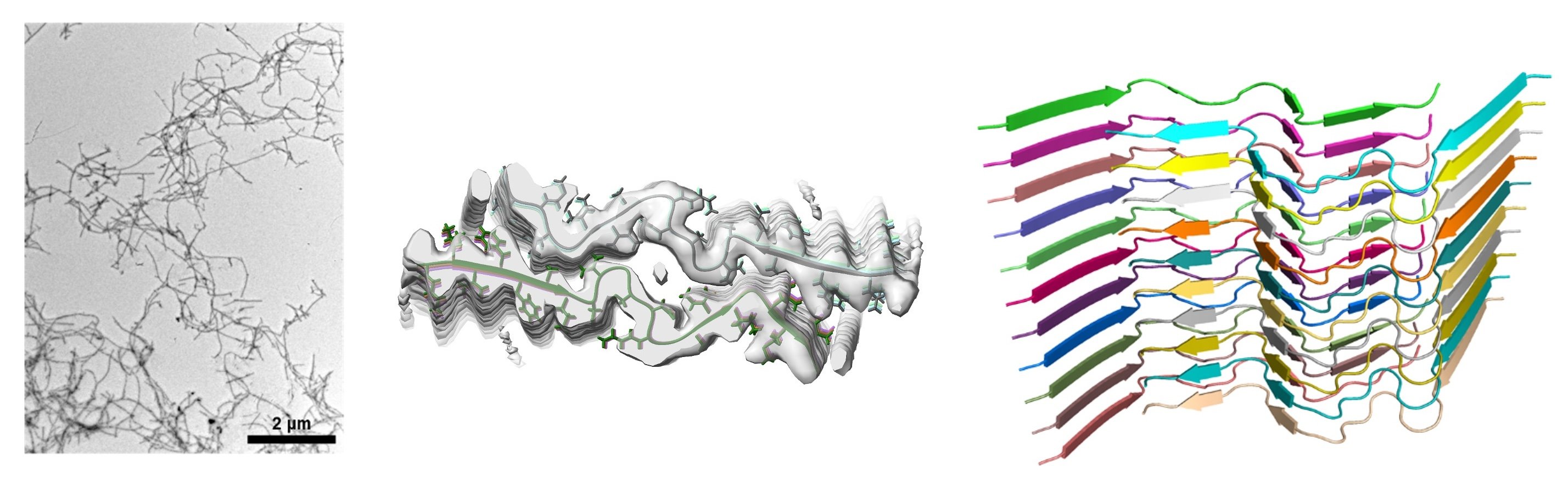
Fig. 2
Structure of Tottori-type amyloid fibrils. Outer shape of amyloid fibrils (left), cross-section of fibrils (center), and orientation of amyloid peptides (right).